1
2
3
4
5
6
7
8
9
10
11
12
13
14
15
16
17
18
19
20
21
22
23
24
25
26
27
28
29
30
31
32
33
34
35
36
37
38
39
40
41
42
43
44
45
46
47
48
49
50
51
52
53
54
55
56
57
58
59
60
61
62
63
64
65
66
67
68
69
70
71
72
73
74
75
76
77
78
79
80
81
82
83
84
85
86
87
88
89
90
91
92
93
94
95
96
97
98
99
100
101
102
103
104
105
106
107
108
109
110
111
112
113
114
115
116
117
118
119
120
121
122
123
124
125
126
127
128
129
130
131
132
133
134
135
136
137
138
139
140
141
142
143
144
145
146
147
|
from builtins import range
from classifier.svm_classifier import LinearSVM
import pandas as pd
import numpy as np
import math
from sklearn import utils
from sklearn.model_selection import train_test_split
import matplotlib.pyplot as plt
import warnings
warnings.filterwarnings("ignore")
def load_iris(iris_path, shuffle=True, tsize=0.8):
"""
加载iris数据
"""
data = pd.read_csv(iris_path, header=0, delimiter=',')
if shuffle:
data = utils.shuffle(data)
species_dict = {
'Iris-setosa': 0,
'Iris-versicolor': 1,
'Iris-virginica': 2
}
data['Species'] = data['Species'].map(species_dict)
data_x = np.array(
[data['SepalLengthCm'], data['SepalWidthCm'], data['PetalLengthCm'], data['PetalWidthCm']]).T
data_y = data['Species']
x_train, x_test, y_train, y_test = train_test_split(data_x, data_y, train_size=tsize, test_size=(1 - tsize),
shuffle=False)
return np.array(x_train), np.array(x_test), np.array(y_train), np.array(y_test)
def load_german_data(data_path, shuffle=True, tsize=0.8):
data_list = pd.read_csv(data_path, header=None, sep='\s+')
data_array = data_list.values
height, width = data_array.shape[:2]
data_x = data_array[:, :(width - 1)]
data_y = data_array[:, (width - 1)]
x_train, x_test, y_train, y_test = train_test_split(data_x, data_y, train_size=tsize, test_size=(1 - tsize),
shuffle=shuffle)
y_train = np.array(list(map(lambda x: 1 if x == 2 else 0, y_train)))
y_test = np.array(list(map(lambda x: 1 if x == 2 else 0, y_test)))
return x_train, x_test, y_train, y_test
def compute_accuracy(y, y_pred):
num = y.shape[0]
num_correct = np.sum(y_pred == y)
acc = float(num_correct) / num
return acc
def cross_validation(x_train, y_train, x_val, y_val, lr_choices, reg_choices):
results = {}
best_val = -1
best_svm = None
for lr in lr_choices:
for reg in reg_choices:
svm = LinearSVM()
svm.train(x_train, y_train, learning_rate=lr, reg=reg, num_iters=2000, batch_size=100, verbose=True)
y_train_pred = svm.predict(x_train)
y_val_pred = svm.predict(x_val)
train_acc = np.mean(y_train_pred == y_train)
val_acc = np.mean(y_val_pred == y_val)
results[(lr, reg)] = (train_acc, val_acc)
if best_val < val_acc:
best_val = val_acc
best_svm = svm
return results, best_svm, best_val
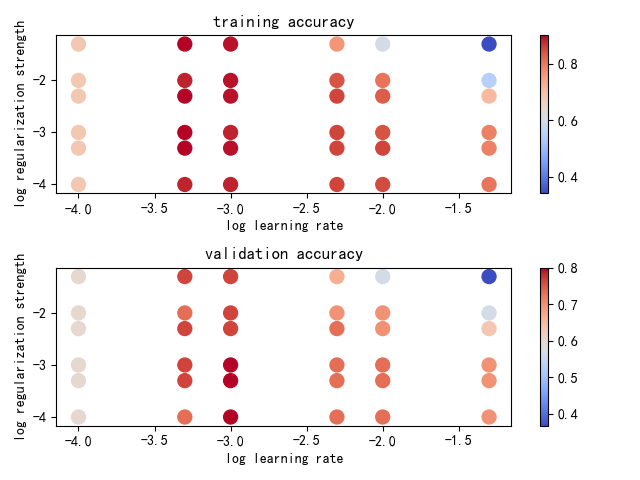
def plot(results):
x_scatter = [math.log10(x[0]) for x in results]
y_scatter = [math.log10(x[1]) for x in results]
marker_size = 100
colors = [results[x][0] for x in results]
plt.subplot(2, 1, 1)
plt.scatter(x_scatter, y_scatter, marker_size, c=colors, cmap=plt.cm.coolwarm)
plt.colorbar()
plt.xlabel('log learning rate')
plt.ylabel('log regularization strength')
plt.title('training accuracy')
colors = [results[x][1] for x in results]
plt.subplot(2, 1, 2)
plt.scatter(x_scatter, y_scatter, marker_size, c=colors, cmap=plt.cm.coolwarm)
plt.colorbar()
plt.xlabel('log learning rate')
plt.ylabel('log regularization strength')
plt.title('validation accuracy')
plt.show()
if __name__ == '__main__':
iris_path = '/home/zj/data/iris-species/Iris.csv'
x_train, x_test, y_train, y_test = load_iris(iris_path, shuffle=True, tsize=0.8)
x_train = x_train.astype(np.double)
x_test = x_test.astype(np.double)
mu = np.mean(x_train, axis=0)
var = np.var(x_train, axis=0)
eps = 1e-8
x_train = (x_train - mu) / np.sqrt(var + eps)
x_test = (x_test - mu) / np.sqrt(var + eps)
lr_choices = [1e-4, 5e-4, 1e-3, 5e-3, 1e-2, 5e-2]
reg_choices = [1e-4, 5e-4, 1e-3, 5e-3, 1e-2, 5e-2]
results, best_svm, best_val = cross_validation(x_train, y_train, x_test, y_test, lr_choices, reg_choices)
plot(results)
for k in results.keys():
lr, reg = k
train_acc, val_acc = results[k]
print('lr = %f, reg = %f, train_acc = %f, val_acc = %f' % (lr, reg, train_acc, val_acc))
print('最好的设置是: lr = %f, reg = %f' % (best_svm.lr, best_svm.reg))
print('最好的测试精度: %f' % best_val)
|
未找到相关的 Issues 进行评论
请联系 @zjykzj 初始化创建